CERoPath Barcoding Tool RodentSEA
RodentSEA
Visit RodentSEA Identification Tool
In order to facilitate the work of field mammalogists in South East Asia, CERoPath developed a reference Barcoding database (RDBSEA), which is available online and allows species assignation by reference of a DNA database (RodentSEA ).
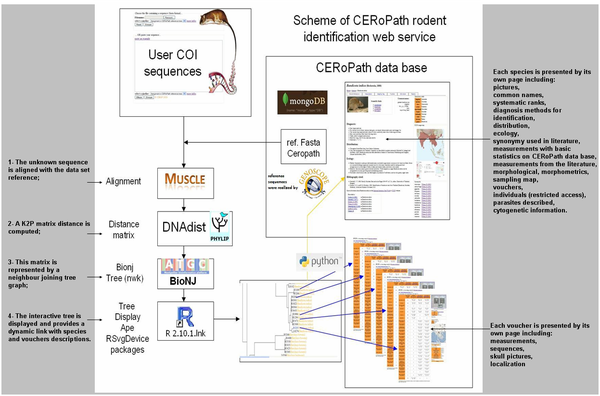
RodentSEA accepts DNA sequences from the barcode region and returns a taxonomic assignment to the species level when possible. It offers a reliable expert system for systematic assessment between an unknown sequence submitted by the user and a collection of reference sequences. This comparison is based on K2P (Kimura 2 Parameter) distances matrix computation and tree representation using Neighbour -Joining clustering method (Fig.1).
Figure
1
: Structure of
RodentSEA
identification tool and the links with the virtual museum
to present vouchers and species information.
Guideline for COI amplification under standard PCR conditions:
Table 4 : Generalist primers used to obtain amplicons of COI genes and produce references sequences (See the list of taxa assessed by the CERoPath barcoding tool below)
| Gene | Type of primer | Name | Primer | |||
| COI gene | generalist | BatL5310 | 5’ CCTACTCRGCCATTTTACCTATG 3’ | |||
| generalist | R6036R | 5’ ACTTCTGGGTGTCCAAAGAATCA 3’ |
1. Perform PCR amplifications in 35µl reaction volumes with 3,5 µl of DNA matrix pre-diluted forty times after extraction (if using Qiagen Spin-Column Protocol).
2. Add 0,7µl for other primer pairs BatL5310 and R6036R (10µM) (Table 4), 4,2µl of a desoxynucleoside-triphosphate mixture (dNTP 2,5 mM), 3,5µl of reaction red Coral Load buffer (10*) (Qiagen), 0,7µl of MgCl2 (25mM) and 0,28 µl of taq DNA polymerase (Qiagen).
3. Use the following thermal cycling parameters: 4 min at 94°C, 40 cycles (30 s at 94°C, 30 s at 48°C, 60 s at 72°C), with a final extension of 10 min at 72°C.
List of taxa assessed by the CERoPath barcoding tool:
Bandicota indica (Bechstein, 1800), Bandicota savilei (Thomas, 1916), Berylmys berdmorei (Blyth, 1851), Berylmys bowersi (Anderson, 1879), Cannomys badius (Hodgson, 1841), Chiropodomys gliroides (Blyth, 1856), Hapalomys delacouri (Thomas, 1927), Leopoldamys edwardsi (Thomas, 1882), Leopoldamys neilli (Marshall, 1976), Leopoldamys sabanus (Thomas, 1887), Maxomys surifer (Miller, 1900), Mus caroli (Bonhote, 1902), Mus cervicolor (Hodgson, 1845), Mus cookii (Ryley, 1914), Mus fragilicauda (Auffray et al., 2003), Niviventer fulvescens (Gray, 1847), Niviventer hinpoon (Marshall, 1976), Niviventer langbianis (Robinson and Kloss, 1922), Niviventer tenaster (Thomas, 1916), Rattus andamanensis (Blyth, 1860), Rattus argentiventer (Robinson and Kloss,1916), Rattus exulans (Peale, 1848), Rattus losea (Swinhoe, 1870), Rattus nitidus (Hodgson, 1845), Rattus norvegicus (Berkenhout, 1769), Rattus phylogenetic r3 (taxonomic statute unclear), Rattus tanezumi (Temminck, 1844), Rattus tiomanicus (Miller, 1900), Rhizomys pruinosus (Blyth, 1851).
|
Guideline for using the RodentSEA :
Perform COI sequences for the samples to be assigned at the species level.
1- Connect to the online CERoPath Barcoding tool:
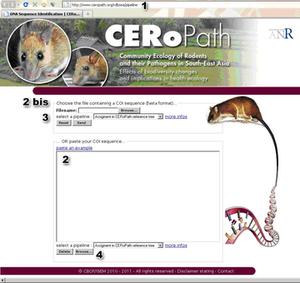
2- Either paste the COI sequence on the dedicated window (Fig 2, nb 2) or upload a file containing COI sequences in fasta format (Fig 2, nb 2bis).
3- Select the reference dataset (Fig 2, nb 3), either the CERoPath dataset, or the CERoPath dataset with additional data from the literature made available online.
4- Send the query (Fig 2 nb 4).
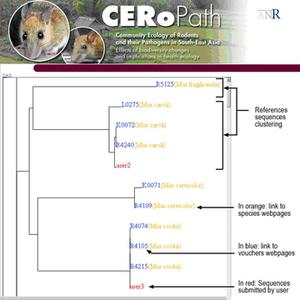
5- The analysis returns a phylogenetic tree, which places the unknown data sequence among the reference samples of the database (Fig. 3).
Detailed information about species and vouchers in the reference database can be accessed from the website. In 2010, the reference database of CERoPath was constituted by 170 individuals belonging to 27 species sampled in Cambodia, Laos and Thailand. 156 of these individuals were used in two taxonomic issues of the CERoPath project (Badenhorst, et al. 2009, Pagès, et al. 2010). The number of species and specimens of the database might increase after 2010. |
Figure 2 : RodentSEA identification tool based on COI comparison between a user sequence and the CERoPath data set reference
|
|
Figure 3 : Interactive tree generated by RodentSEA identification tool after alignment of unknown sequences |
The RodentSEA is connected to a large rodent database (Rodent information center ) that can be consulted online and present molecular, morphometrical, museological, parasitological, cytogenetical, phylogenetical, ecological and spatial information collected by the project and ensure that these data are clearly linked to vouchers “specimen”.
© Cirad 2013 - All rights reserved - Disclaimer stating - Contact